Pathomics
Pathomics refers to the high‑throughput quantitative analysis of digital histopathology images, typically whole‑slide images, to extract measurable information about tissue architecture, cell morphology, texture, and spatial organization (Figure 1). By transforming placental histology into structured quantitative data, pathomics complements conventional microscopy and enables more reproducible and scalable assessment of placental structure and disease. This is particularly valuable for the placenta, an organ characterized by marked spatial heterogeneity and dynamic changes across gestation.

Recent placental applications illustrate this potential: deep learning models such as GestAltNet have estimated gestational age from placental whole‑slide images; hierarchical convolutional approaches have identified decidual vasculopathy; whole‑slide learning models have classified villous infarction and other parenchymal lesions; and computational pathology models have distinguished preeclamptic placentas from controls.

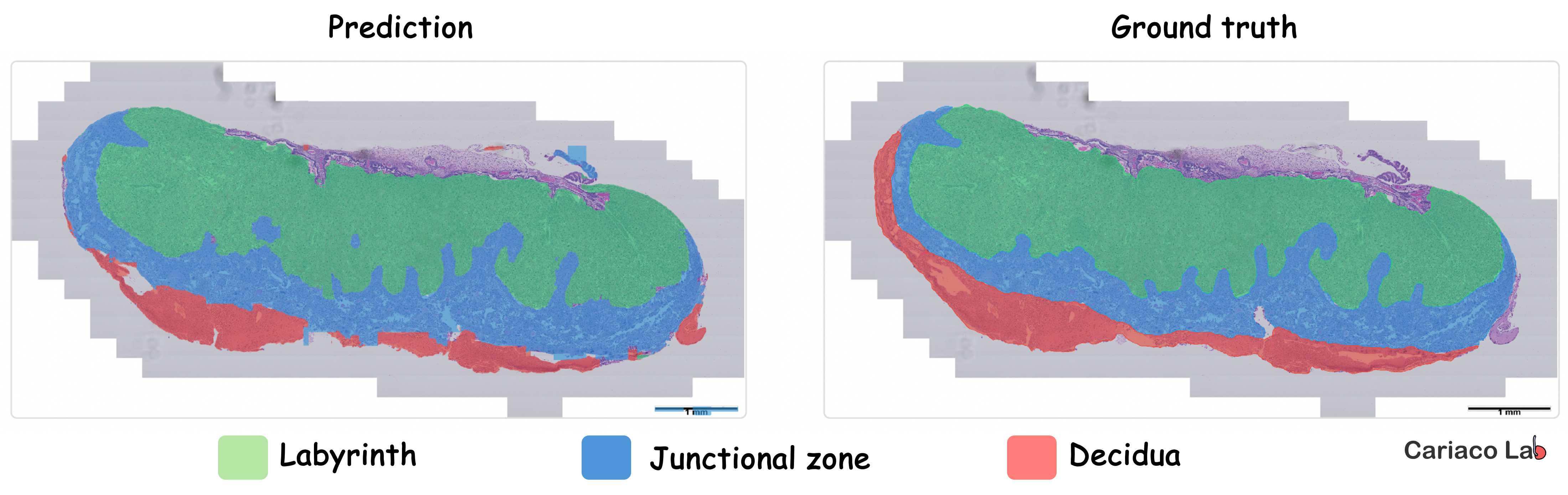
Notably, the HAPPY pipeline (Histology Analysis Pipeline.PY) provides a biologically interpretable framework for quantifying nuclei, cell types, and tissue microstructures across placenta whole‑slide images, and subsequent work has shown that lesion‑associated changes can extend beyond visibly abnormal regions. We are now applying this framework in our laboratory to generate quantitative whole-slide human placental phenotypes in our own cohorts (Figure 2), while also developing AI‑powered image analysis and deep learning workflows to segment tissue regions, classify cell types, and quantify their spatial organization in mouse placentas (Figure 3).
Together, these studies suggest that placental pathomics can improve diagnostic reproducibility, reveal subtle disease‑associated morphology, and support biomarker discovery, although broader validation and standardization are still needed for routine clinical implementation.
References- Bülow RD, Jäger PF, Isensee F, et al. Extending the landscape of omics technologies by pathomics. 2023.
- Hosseini MS, Bera K, Wang S, et al. Computational pathology: A survey review and the way forward. 2024.
- Mobadersany P, Cooper LAD, Goldstein JA. GestAltNet: aggregation and attention to improve deep learning of gestational age from placental whole-slide images. 2021.
- Clymer D, Jimenez JA, Mamoon A, et al. Decidual vasculopathy identification in whole slide images using multiresolution hierarchical convolutional neural networks. 2020.
- Vanea C, Campbell J, Dodi O, et al. Mapping cell-to-tissue graphs across human placenta histology whole slide images using deep learning with HAPPY. 2024.
- Walker EC, Vanea C, Ernst LM, et al. Biologically Inspired Digital Histology for Deep Phenotyping of Placental Composition Changes Across Major Lesion Types. 2025.
- d’Amati A, Baldini GM, Difonzo T, et al. Artificial Intelligence in Placental Pathology: New Diagnostic Imaging Tools in Evolution and in Perspective. 2025.